Publication in Nature Medicine:
Large genomic study on perivascular space burden unravels early mechanisms of cerebral small vessel disease
Retour

The results of a large genomic study on perivascular spaces, coordinated by researchers from the ELEANORE team and the Institute of Clinical Neurodegenerative Diseases at the University Hospital of Bordeaux, have been published in the journal Nature Medicine.
The study reveals mechanisms involved early in the disease of small cerebral arteries.
Cerebral small vessel disease is an age-related disease that results from an alteration in the structure and/or function of the small arteries that supply blood to the brain.
It is a leading cause of stroke and dementia for which no mechanism-based drugs are currently available.
Among the cerebral lesions that characterize it are the perivascular space burden (PVS) detectable on cerebral MRI.
These spaces filled with biological fluid surround the blood vessels for a short distance as they enter the brain.
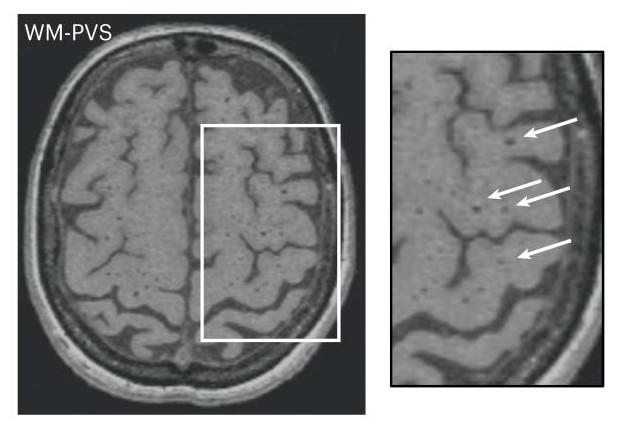
loading in the white matter
The first genomic study on PVS revealed 24 genetic risk loci for PVS in a large meta-analysis of genome-wide association studies on more than 40,000 participants. The majority of these regions, identified in people aged 65 years on average and mainly of European origin, were also associated with PVS in two smaller studies, including young adults in their 20s (i-Share study) and participants from East Asia.
This study identified that a large number of PVS-associated genes are involved in brain developmental diseases and expressed in brain vascular cells before birth, suggesting a link between this marker of small brain artery disease and developmental processes.
Bioinformatics analysis reveal 12 genes to prioritize for functional follow-up, including targets of drugs investigated for vascular and cognitive disorders. These findings provide entirely novel insights into the biology and clinical significance of PVS and open new avenues for therapeutic developments.
says Stephanie Debette, Professor of Epidemiology and Neurologist at the University of Bordeaux and Bordeaux University Hospital, and Director of the Bordeaux Population Health research center (Inserm / University of Bordeaux), one of the corresponding authors of the current study.
comments Marie-Gabrielle Duperron, Inserm-Bettencourt fellow researcher at University of Bordeaux, Inserm, and Bordeaux University Hospital, first author of the study.
These results of the first genomic study on EPV were published, April 17, 2023, in the journal Nature Medicine. The study was based on DNA samples from more than 40,000 participants from numerous general population cohorts and biobanks around the world, participating in different international scientific consortia (CHARGE, BRIDGET and ISGC).
It has been made possible by numerous research funds, including the RHU SHIVA, which benefits from a government grant managed by the Agence Nationale de la Recherche (ANR) under the future investment program.
This collaborative project was coordinated by researchers from the Bordeaux Population Health (University of Bordeaux and Inserm U1219) and the Neurodegeneratives diseases institute of Bordeaux. It was co-coordinated by researchers from the Latin American Brain Health (BrainLat), University of Santiago (Chile) and Erasmus MC University Medical Center in Rotterdam (The Netherlands), in close collaboration with research teams from the Biggs Institute, UTHealth San Antonio (USA), the University of Edinburgh (UK) and Kyoto University (Japan).
Link to publication
Genomics of perivascular space burden unravels early mechanisms of cerebral small vessel disease
(2023).
Marie-Gabrielle Duperron, Maria J. Knol, Quentin Le Grand, Tavia E. Evans, Aniket Mishra, Ami
Tsuchida, et al. Nature Medicine. doi : 10.1038/s41591-023-02268-w
Link to research briefing (summary)
https://rdcu.be/c9O2T
More information
Pr. Stéphanie Debette
Université de Bordeaux, Inserm, Centre de recherche en santé des populations de Bordeaux, UMR 1219
Service de neurologie, CHU de Bordeaux
146, rue Léo Saignat, 33076 Bordeaux, France
Courriel : stephanie.debette@u-bordeaux.fr


